IRF3 Protein Hotspots Mapping
MixMD cosolvent simulations for identifying druggable hotspots and therapeutic targets
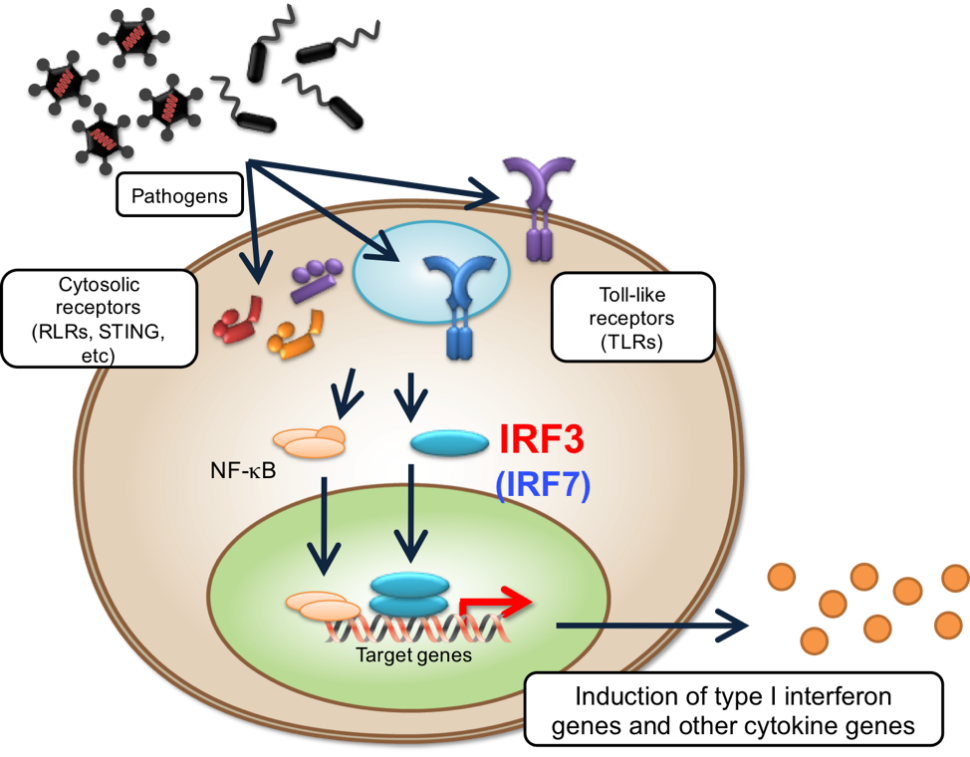
IRF3 signalling pathway
Courtesy of Tadatsugu Taniguchi, Ph.D.
This research focuses on mapping druggable hotspots on the Interferon Regulatory Factor 3 (IRF3) protein surface using advanced MixMD cosolvent molecular dynamics simulations. IRF3 is a crucial transcription factor in the innate immune response pathway, making it an important therapeutic target for immune-related disorders and cancer.
By employing drug-like fragments including Acetonitrile, Ethanol, Isopropanol, Imidazole, and Pyrimidine in cosolvent simulations, we identify and characterize potential binding sites that could serve as targets for small molecule drug development. This computational approach provides valuable insights for structure-based drug design efforts.
In addition, these simulations enable the discovery of allosteric binding pockets that may remain hidden in experimental structures, offering new opportunities to modulate IRF3 function with high specificity. This research lays the groundwork for designing novel therapeutics to control dysregulated immune signaling and develop treatments for autoimmune diseases, viral infections, and cancer.
MixMD Cosolvent Simulations
MixMD Cosolvent Simulations provide a powerful approach to mapping druggable hotspots on protein surfaces by tracking the behavior of probe or cosolvent molecules in molecular dynamics simulations. By immersing a protein in a mixture of water and small organic probes, MixMD identifies regions where these probes preferentially accumulate, highlighting pockets that are energetically favorable for ligand binding. These simulations go beyond static structural analysis by capturing dynamic protein–solvent interactions, offering insights into hidden or transient binding sites that are often missed in traditional methods. This makes MixMD a valuable tool in rational drug discovery, guiding the design of small molecules that can effectively target proteins with high specificity and affinity.